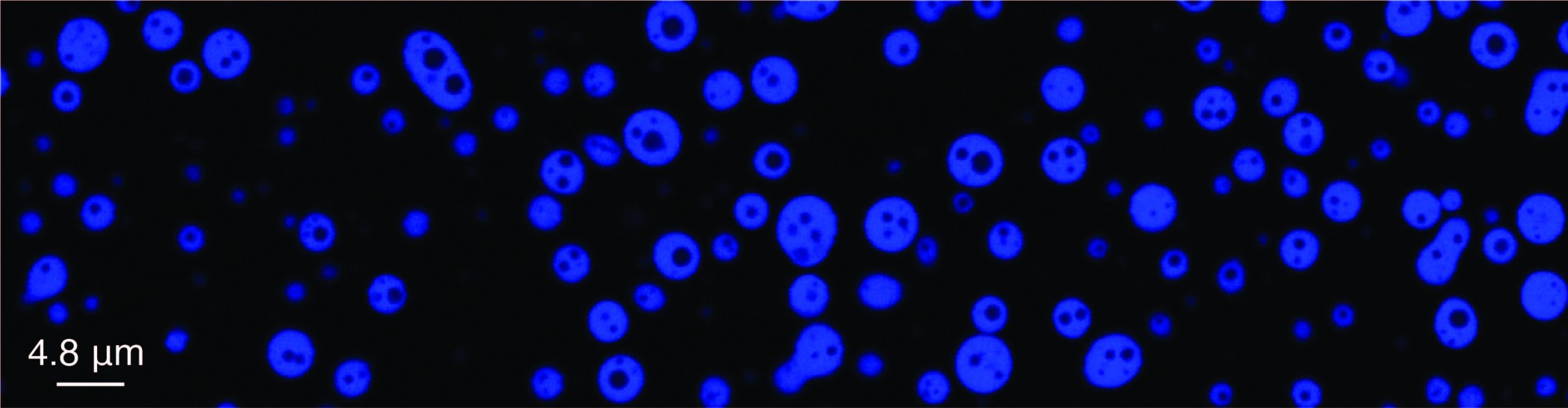
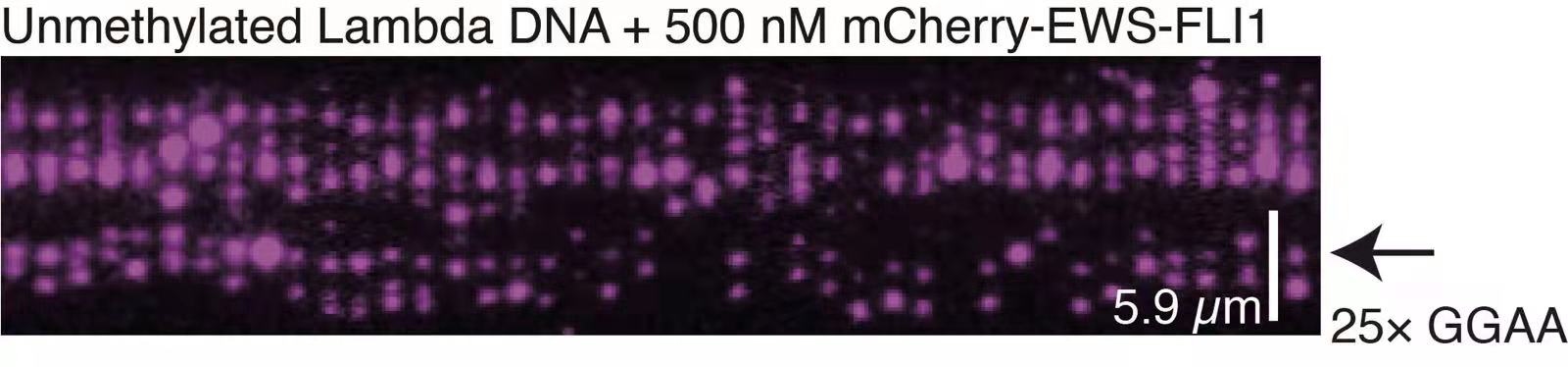
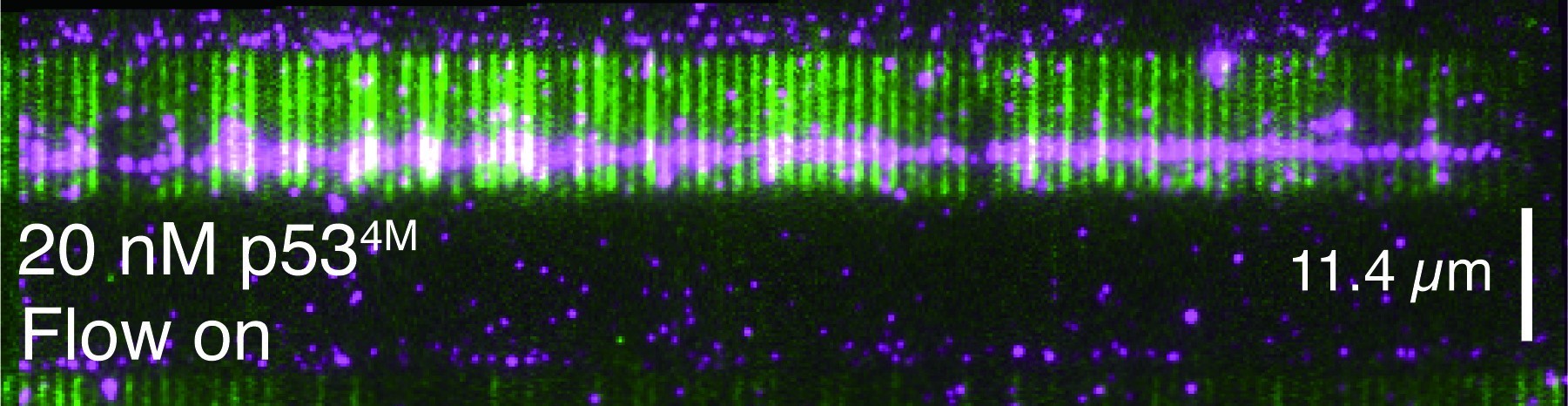
The ultimate goal of research in my independent laboratory is to develop cutting-edge single-molecule biophysical technologies and establish non-equilibrium physics models to understand the molecular mechanisms underlying functional biomolecular dynamic clusters in molecular biology. Our research focuses on two types of biomolecular dynamic clusters: dsDNA-protein co-condensation and ssDNA-RPA complex. These studies have the potential to lead to the development of new strategies for intervening in human diseases.
Advertisements and feature films of Zhi QI Lab:
https://www.bilibili.com/video/BV1wk4y1m75Q